The findings of a large-scale screen could help researchers design nanoparticles that target specific types of cancer.
Anne Trafton | MIT News Office
Publication Date: July 21, 2022
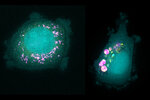
MIT researchers have identified biomarkers that predict whether different types of cancer cells will take up specific nanoparticles. In this image, cells with low levels of a protein called SLC46A3 (left) take up particles called liposomes (pink), while cells with high levels of SLC46A3 (right) do not. Credits: Image: Courtesy of the researchers
Using nanoparticles to deliver cancer drugs offers a way to hit tumors with large doses of drugs while avoiding the harmful side effects that often come with chemotherapy. However, so far, only a handful of nanoparticle-based cancer drugs have been FDA-approved.
A new study from MIT and Broad Institute of MIT and Harvard researchers may help to overcome some of the obstacles to the development of nanoparticle-based drugs. The team’s analysis of the interactions between 35 different types of nanoparticles and nearly 500 types of cancer cells revealed thousands of biological traits that influence whether those cells take up different types of nanoparticles.
The findings could help researchers better tailor their drug-delivery particles to specific types of cancer, or design new particles that take advantage of the biological features of particular types of cancer cells.
“We are excited by our findings because it is really just the beginning — we can use this approach to map out what types of nanoparticles are best to target certain cell types, from cancer to immune cells and other kinds of healthy and diseased organ cells. We are learning how surface chemistry and other material properties play a role in targeting,” says Paula Hammond, an MIT Institute Professor, head of the Department of Chemical Engineering, and a member of MIT’s Koch Institute for Integrative Cancer Research.
Hammond is the senior author of the new study, which appears today in Science. The paper’s lead authors are Natalie Boehnke, an MIT postdoc who will soon join the faculty at the University of Minnesota, and Joelle Straehla, the Charles W. and Jennifer C. Johnson Clinical Investigator at the Koch Institute, an instructor at Harvard Medical School, and a pediatric oncologist at Dana-Farber Cancer Institute.
Hammond’s lab has previously developed many types of nanoparticles that can be used to deliver drugs to cells. Studies in her lab and others have shown that different types of cancer cells often respond differently to the same nanoparticles. Boehnke, who was studying ovarian cancer when she joined Hammond’s lab, and Straehla, who was studying brain cancer, also noticed this phenomenon in their studies.
The researchers hypothesized that biological differences between cells could be driving the variation in their responses. To figure out what those differences might be, they decided to pursue a large-scale study in which they could look at a huge number of different cells interacting with many types of nanoparticles.
Straehla had recently learned about the Broad Institute’s PRISM platform, which was designed to allow researchers to rapidly screen thousands of drugs on hundreds of different cancer types at the same time. With instrumental collaboration from Angela Koehler, an MIT associate professor of biological engineering, the team decided to try to adapt that platform to screen cell-nanoparticle interactions instead of cell-drug interactions.
“Using this approach, we can start thinking about whether there is something about a cell’s genotypic signature that predicts how many nanoparticles it will take up,” Boehnke says.
For their screen, the researchers used 488 cancer cell lines from 22 different tissues of origin. Each cell type is “barcoded” with a unique DNA sequence that allows researchers to identify the cells later on. For each cell type, extensive datasets are also available on their gene expression profiles and other biological characteristics.
On the nanoparticle side, the researchers created 35 particles, each of which had a core consisting of either liposomes (particles made from many fatty molecules called lipids), a polymer known as PLGA, or another polymer called polystyrene. The researchers also coated the particles with different types of protective or targeting molecules, including polymers such as polyethylene glycol, antibodies, and polysaccharides. This allowed them to study the influence of both the core composition and the surface chemistry of the particles.
Working with Broad Institute scientists, including Jennifer Roth, director of the PRISM lab, the researchers exposed pools of hundreds of different cells to one of 35 different nanoparticles. Each nanoparticle had a fluorescent tag, so the researchers could use a cell-sorting technique to separate the cells based on how much fluorescence they gave off after an exposure of either four or 24 hours.
Based on these measurements, each cell line was assigned a score representing its affinity for each nanoparticle. The researchers then used machine learning algorithms to analyze those scores along with all of the other biological data available for each cell line.
This analysis yielded thousands of features, or biomarkers, associated with affinity for different types of nanoparticles. Many of these markers were genes that code for the cellular machinery needed to bind particles, bring them into a cell, or process them. Some of these genes were already known to be involved in nanoparticle trafficking, but many others were new.
“We found some markers that we expected, and we also found much more that has really been unexplored. We're hoping that other people can use this dataset to help expand their view of how nanoparticles and cells interact,” Straehla says.
The researchers picked out one of the biomarkers they identified, a protein called SLC46A3, for further study. The PRISM screen had shown that high levels of this protein correlated with very low uptake of lipid-based nanoparticles. When the researchers tested these particles in mouse models of melanoma, they found the same correlation. The findings suggest that this biomarker could be used to help doctors identify patients whose tumors are more likely to respond to nanoparticle-based therapies.
Now, the researchers are trying to uncover the mechanism of how SLC46A3 regulates nanoparticle uptake. If they could discover new ways to decrease cellular levels of this protein, that could help make tumors more susceptible to drugs carried by lipid nanoparticles. The researchers are also working on further exploring some of the other biomarkers they found.
This screening approach could also be used to investigate many other types of nanoparticles that the researchers didn’t look at in this study.
“The sky is the limit in terms of what other undiscovered biomarkers are out there that we just haven't captured because we haven't screened them,” Boehnke says. “Hopefully it’s an inspiration for others to start looking at their nanoparticle systems in a similar manner.”
The research was funded, in part, by SPARC funding to the Broad Institute, the Marble Center for Cancer Nanomedicine at the Koch Institute, and the Koch Institute Support (core) Grant from the National Cancer Institute.
RELATED LINKS
By adapting the MIT Broad Institute’s PRISM platform, which was designed to allow researchers to rapidly screen thousands of drugs on hundreds of different cancer types at the same time, the authors were able to track nanoparticles and the chemicals they contained and observe their efficacy across many strains of cancer and to validate the biomarkers. Their work establishes the power of integrated screens for nanoparticle delivery and enables the identification and utilization of biomarkers to rationally design nanoformulations.
Hartmann352
Anne Trafton | MIT News Office
Publication Date: July 21, 2022

MIT researchers have identified biomarkers that predict whether different types of cancer cells will take up specific nanoparticles. In this image, cells with low levels of a protein called SLC46A3 (left) take up particles called liposomes (pink), while cells with high levels of SLC46A3 (right) do not. Credits: Image: Courtesy of the researchers
Using nanoparticles to deliver cancer drugs offers a way to hit tumors with large doses of drugs while avoiding the harmful side effects that often come with chemotherapy. However, so far, only a handful of nanoparticle-based cancer drugs have been FDA-approved.
A new study from MIT and Broad Institute of MIT and Harvard researchers may help to overcome some of the obstacles to the development of nanoparticle-based drugs. The team’s analysis of the interactions between 35 different types of nanoparticles and nearly 500 types of cancer cells revealed thousands of biological traits that influence whether those cells take up different types of nanoparticles.
The findings could help researchers better tailor their drug-delivery particles to specific types of cancer, or design new particles that take advantage of the biological features of particular types of cancer cells.
“We are excited by our findings because it is really just the beginning — we can use this approach to map out what types of nanoparticles are best to target certain cell types, from cancer to immune cells and other kinds of healthy and diseased organ cells. We are learning how surface chemistry and other material properties play a role in targeting,” says Paula Hammond, an MIT Institute Professor, head of the Department of Chemical Engineering, and a member of MIT’s Koch Institute for Integrative Cancer Research.
Hammond is the senior author of the new study, which appears today in Science. The paper’s lead authors are Natalie Boehnke, an MIT postdoc who will soon join the faculty at the University of Minnesota, and Joelle Straehla, the Charles W. and Jennifer C. Johnson Clinical Investigator at the Koch Institute, an instructor at Harvard Medical School, and a pediatric oncologist at Dana-Farber Cancer Institute.
Hammond’s lab has previously developed many types of nanoparticles that can be used to deliver drugs to cells. Studies in her lab and others have shown that different types of cancer cells often respond differently to the same nanoparticles. Boehnke, who was studying ovarian cancer when she joined Hammond’s lab, and Straehla, who was studying brain cancer, also noticed this phenomenon in their studies.
The researchers hypothesized that biological differences between cells could be driving the variation in their responses. To figure out what those differences might be, they decided to pursue a large-scale study in which they could look at a huge number of different cells interacting with many types of nanoparticles.
Straehla had recently learned about the Broad Institute’s PRISM platform, which was designed to allow researchers to rapidly screen thousands of drugs on hundreds of different cancer types at the same time. With instrumental collaboration from Angela Koehler, an MIT associate professor of biological engineering, the team decided to try to adapt that platform to screen cell-nanoparticle interactions instead of cell-drug interactions.
“Using this approach, we can start thinking about whether there is something about a cell’s genotypic signature that predicts how many nanoparticles it will take up,” Boehnke says.
For their screen, the researchers used 488 cancer cell lines from 22 different tissues of origin. Each cell type is “barcoded” with a unique DNA sequence that allows researchers to identify the cells later on. For each cell type, extensive datasets are also available on their gene expression profiles and other biological characteristics.
On the nanoparticle side, the researchers created 35 particles, each of which had a core consisting of either liposomes (particles made from many fatty molecules called lipids), a polymer known as PLGA, or another polymer called polystyrene. The researchers also coated the particles with different types of protective or targeting molecules, including polymers such as polyethylene glycol, antibodies, and polysaccharides. This allowed them to study the influence of both the core composition and the surface chemistry of the particles.
Working with Broad Institute scientists, including Jennifer Roth, director of the PRISM lab, the researchers exposed pools of hundreds of different cells to one of 35 different nanoparticles. Each nanoparticle had a fluorescent tag, so the researchers could use a cell-sorting technique to separate the cells based on how much fluorescence they gave off after an exposure of either four or 24 hours.
Based on these measurements, each cell line was assigned a score representing its affinity for each nanoparticle. The researchers then used machine learning algorithms to analyze those scores along with all of the other biological data available for each cell line.
This analysis yielded thousands of features, or biomarkers, associated with affinity for different types of nanoparticles. Many of these markers were genes that code for the cellular machinery needed to bind particles, bring them into a cell, or process them. Some of these genes were already known to be involved in nanoparticle trafficking, but many others were new.
“We found some markers that we expected, and we also found much more that has really been unexplored. We're hoping that other people can use this dataset to help expand their view of how nanoparticles and cells interact,” Straehla says.
The researchers picked out one of the biomarkers they identified, a protein called SLC46A3, for further study. The PRISM screen had shown that high levels of this protein correlated with very low uptake of lipid-based nanoparticles. When the researchers tested these particles in mouse models of melanoma, they found the same correlation. The findings suggest that this biomarker could be used to help doctors identify patients whose tumors are more likely to respond to nanoparticle-based therapies.
Now, the researchers are trying to uncover the mechanism of how SLC46A3 regulates nanoparticle uptake. If they could discover new ways to decrease cellular levels of this protein, that could help make tumors more susceptible to drugs carried by lipid nanoparticles. The researchers are also working on further exploring some of the other biomarkers they found.
This screening approach could also be used to investigate many other types of nanoparticles that the researchers didn’t look at in this study.
“The sky is the limit in terms of what other undiscovered biomarkers are out there that we just haven't captured because we haven't screened them,” Boehnke says. “Hopefully it’s an inspiration for others to start looking at their nanoparticle systems in a similar manner.”
The research was funded, in part, by SPARC funding to the Broad Institute, the Marble Center for Cancer Nanomedicine at the Koch Institute, and the Koch Institute Support (core) Grant from the National Cancer Institute.
RELATED LINKS
- Paula Hammond
- Natalie Boehnke
- Joelle Straehla
- Koch Institute
- PRISM Lab
- Department of Chemical Engineering
- School of Engineering
- Broad Institute
By adapting the MIT Broad Institute’s PRISM platform, which was designed to allow researchers to rapidly screen thousands of drugs on hundreds of different cancer types at the same time, the authors were able to track nanoparticles and the chemicals they contained and observe their efficacy across many strains of cancer and to validate the biomarkers. Their work establishes the power of integrated screens for nanoparticle delivery and enables the identification and utilization of biomarkers to rationally design nanoformulations.
Hartmann352


